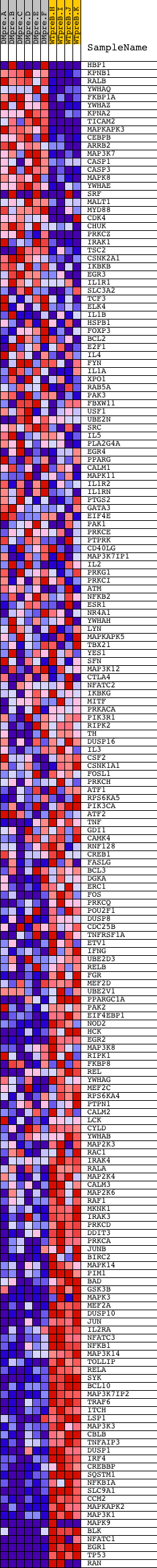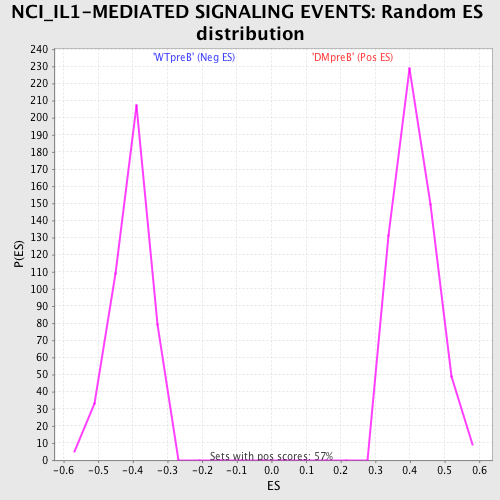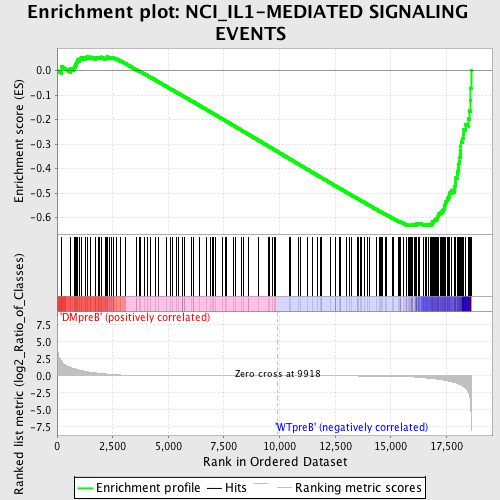
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_DMpreB_versus_WTpreB.phenotype_DMpreB_versus_WTpreB.cls #DMpreB_versus_WTpreB |
| Phenotype | phenotype_DMpreB_versus_WTpreB.cls#DMpreB_versus_WTpreB |
| Upregulated in class | WTpreB |
| GeneSet | NCI_IL1-MEDIATED SIGNALING EVENTS |
| Enrichment Score (ES) | -0.6337172 |
| Normalized Enrichment Score (NES) | -1.5677052 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.06668101 |
| FWER p-Value | 0.691 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | HBP1 | 2048 2162 21086 | 207 | 2.101 | 0.0158 | No | ||
| 2 | KPNB1 | 20274 | 617 | 1.219 | 0.0094 | No | ||
| 3 | RALB | 13857 | 760 | 1.049 | 0.0152 | No | ||
| 4 | YWHAQ | 10369 | 828 | 0.997 | 0.0244 | No | ||
| 5 | FKBP1A | 2801 | 880 | 0.937 | 0.0337 | No | ||
| 6 | YWHAZ | 10370 | 915 | 0.915 | 0.0436 | No | ||
| 7 | KPNA2 | 9245 | 1026 | 0.826 | 0.0483 | No | ||
| 8 | TICAM2 | 23440 | 1095 | 0.778 | 0.0546 | No | ||
| 9 | MAPKAPK3 | 19003 | 1265 | 0.664 | 0.0540 | No | ||
| 10 | CEBPB | 8733 | 1369 | 0.609 | 0.0563 | No | ||
| 11 | ARRB2 | 20806 | 1508 | 0.540 | 0.0558 | No | ||
| 12 | MAP3K7 | 16255 | 1728 | 0.446 | 0.0496 | No | ||
| 13 | CASP1 | 19577 | 1743 | 0.440 | 0.0545 | No | ||
| 14 | CASP3 | 8693 | 1868 | 0.395 | 0.0529 | No | ||
| 15 | MAPK8 | 6459 | 1918 | 0.379 | 0.0551 | No | ||
| 16 | YWHAE | 20776 | 1991 | 0.354 | 0.0558 | No | ||
| 17 | SRF | 22961 1597 | 2181 | 0.296 | 0.0493 | No | ||
| 18 | MALT1 | 6274 | 2207 | 0.289 | 0.0517 | No | ||
| 19 | MYD88 | 18970 | 2257 | 0.274 | 0.0526 | No | ||
| 20 | CDK4 | 3424 19859 | 2258 | 0.274 | 0.0561 | No | ||
| 21 | CHUK | 23665 | 2349 | 0.253 | 0.0545 | No | ||
| 22 | PRKCZ | 5260 | 2458 | 0.222 | 0.0515 | No | ||
| 23 | IRAK1 | 4916 | 2512 | 0.209 | 0.0513 | No | ||
| 24 | TSC2 | 23090 | 2549 | 0.201 | 0.0519 | No | ||
| 25 | CSNK2A1 | 14797 | 2669 | 0.175 | 0.0477 | No | ||
| 26 | IKBKB | 4907 | 2832 | 0.138 | 0.0407 | No | ||
| 27 | EGR3 | 4656 | 3054 | 0.102 | 0.0300 | No | ||
| 28 | IL1R1 | 14264 3987 | 3579 | 0.059 | 0.0024 | No | ||
| 29 | SLC3A2 | 23754 | 3588 | 0.058 | 0.0027 | No | ||
| 30 | TCF3 | 17114 | 3697 | 0.053 | -0.0025 | No | ||
| 31 | ELK4 | 8895 4664 3937 | 3729 | 0.051 | -0.0035 | No | ||
| 32 | IL1B | 14431 | 3930 | 0.043 | -0.0138 | No | ||
| 33 | HSPB1 | 4879 | 4045 | 0.039 | -0.0195 | No | ||
| 34 | FOXP3 | 9804 5426 | 4204 | 0.035 | -0.0276 | No | ||
| 35 | BCL2 | 8651 3928 13864 4435 981 4062 13863 4027 | 4399 | 0.030 | -0.0378 | No | ||
| 36 | E2F1 | 14384 | 4578 | 0.027 | -0.0471 | No | ||
| 37 | IL4 | 9174 | 4896 | 0.022 | -0.0640 | No | ||
| 38 | FYN | 3375 3395 20052 | 5080 | 0.020 | -0.0737 | No | ||
| 39 | IL1A | 4915 | 5169 | 0.019 | -0.0782 | No | ||
| 40 | XPO1 | 4172 | 5368 | 0.017 | -0.0887 | No | ||
| 41 | RAB5A | 11310 | 5448 | 0.016 | -0.0928 | No | ||
| 42 | PAK3 | 9528 | 5625 | 0.015 | -0.1022 | No | ||
| 43 | FBXW11 | 20926 | 5630 | 0.015 | -0.1022 | No | ||
| 44 | USF1 | 5832 10260 10261 | 5657 | 0.015 | -0.1034 | No | ||
| 45 | UBE2N | 8216 19898 | 5725 | 0.014 | -0.1069 | No | ||
| 46 | SRC | 5507 | 5732 | 0.014 | -0.1070 | No | ||
| 47 | IL5 | 20884 10220 | 6024 | 0.012 | -0.1226 | No | ||
| 48 | PLA2G4A | 13809 | 6112 | 0.012 | -0.1272 | No | ||
| 49 | EGR4 | 24483 | 6142 | 0.012 | -0.1286 | No | ||
| 50 | PPARG | 1151 1144 17319 | 6395 | 0.010 | -0.1422 | No | ||
| 51 | CALM1 | 21184 | 6700 | 0.009 | -0.1585 | No | ||
| 52 | MAPK11 | 2264 9618 22163 | 6874 | 0.008 | -0.1678 | No | ||
| 53 | IL1R2 | 14265 | 6984 | 0.008 | -0.1736 | No | ||
| 54 | IL1RN | 15096 | 7041 | 0.008 | -0.1766 | No | ||
| 55 | PTGS2 | 5317 9655 | 7104 | 0.007 | -0.1798 | No | ||
| 56 | GATA3 | 9004 | 7432 | 0.006 | -0.1975 | No | ||
| 57 | EIF4E | 15403 1827 8890 | 7585 | 0.006 | -0.2057 | No | ||
| 58 | PAK1 | 9527 | 7592 | 0.006 | -0.2059 | No | ||
| 59 | PRKCE | 9575 | 7917 | 0.005 | -0.2234 | No | ||
| 60 | PTPRK | 5332 20066 | 8038 | 0.005 | -0.2299 | No | ||
| 61 | CD40LG | 24330 | 8298 | 0.004 | -0.2439 | No | ||
| 62 | MAP3K7IP1 | 2193 2171 22419 | 8369 | 0.004 | -0.2476 | No | ||
| 63 | IL2 | 15354 | 8581 | 0.003 | -0.2591 | No | ||
| 64 | PRKG1 | 5289 | 8607 | 0.003 | -0.2604 | No | ||
| 65 | PRKCI | 9576 | 9032 | 0.002 | -0.2834 | No | ||
| 66 | ATM | 2976 19115 | 9513 | 0.001 | -0.3094 | No | ||
| 67 | NFKB2 | 23810 | 9535 | 0.001 | -0.3105 | No | ||
| 68 | ESR1 | 20097 4685 | 9667 | 0.001 | -0.3176 | No | ||
| 69 | NR4A1 | 9099 | 9752 | 0.000 | -0.3222 | No | ||
| 70 | YWHAH | 5937 10368 | 9813 | 0.000 | -0.3254 | No | ||
| 71 | LYN | 16281 | 10463 | -0.001 | -0.3606 | No | ||
| 72 | MAPKAPK5 | 16387 | 10496 | -0.001 | -0.3623 | No | ||
| 73 | TBX21 | 20276 | 10857 | -0.002 | -0.3818 | No | ||
| 74 | YES1 | 5930 | 10960 | -0.003 | -0.3873 | No | ||
| 75 | SFN | 7060 12062 | 11240 | -0.003 | -0.4024 | No | ||
| 76 | MAP3K12 | 6454 11164 | 11464 | -0.004 | -0.4145 | No | ||
| 77 | CTLA4 | 14238 | 11695 | -0.005 | -0.4269 | No | ||
| 78 | NFATC2 | 5168 2866 | 11816 | -0.005 | -0.4334 | No | ||
| 79 | IKBKG | 2570 2562 4908 | 11885 | -0.005 | -0.4370 | No | ||
| 80 | MITF | 17349 | 11903 | -0.005 | -0.4378 | No | ||
| 81 | PRKACA | 18549 3844 | 12270 | -0.007 | -0.4576 | No | ||
| 82 | PIK3R1 | 3170 | 12501 | -0.008 | -0.4700 | No | ||
| 83 | RIPK2 | 2528 15935 | 12516 | -0.008 | -0.4706 | No | ||
| 84 | TH | 17548 | 12533 | -0.008 | -0.4714 | No | ||
| 85 | DUSP16 | 1004 7699 | 12672 | -0.009 | -0.4788 | No | ||
| 86 | IL3 | 20453 | 12729 | -0.009 | -0.4817 | No | ||
| 87 | CSF2 | 20454 | 12730 | -0.009 | -0.4816 | No | ||
| 88 | CSNK1A1 | 8204 | 13023 | -0.010 | -0.4973 | No | ||
| 89 | FOSL1 | 23779 | 13138 | -0.011 | -0.5034 | No | ||
| 90 | PRKCH | 21246 | 13221 | -0.012 | -0.5077 | No | ||
| 91 | ATF1 | 8634 4417 | 13480 | -0.013 | -0.5215 | No | ||
| 92 | RPS6KA5 | 2076 21005 | 13507 | -0.014 | -0.5227 | No | ||
| 93 | PIK3CA | 9562 | 13536 | -0.014 | -0.5241 | No | ||
| 94 | ATF2 | 4418 2759 | 13658 | -0.015 | -0.5304 | No | ||
| 95 | TNF | 23004 | 13662 | -0.015 | -0.5304 | No | ||
| 96 | GDI1 | 9012 4770 24297 4769 | 13827 | -0.017 | -0.5391 | No | ||
| 97 | CAMK4 | 4473 | 13955 | -0.019 | -0.5457 | No | ||
| 98 | RNF128 | 24236 2641 | 14045 | -0.020 | -0.5503 | No | ||
| 99 | CREB1 | 3990 8782 4558 4093 | 14348 | -0.026 | -0.5664 | No | ||
| 100 | FASLG | 13789 | 14495 | -0.029 | -0.5739 | No | ||
| 101 | BCL3 | 8654 | 14526 | -0.029 | -0.5752 | No | ||
| 102 | DGKA | 3359 19589 | 14560 | -0.030 | -0.5766 | No | ||
| 103 | ERC1 | 1013 17021 995 1136 | 14611 | -0.032 | -0.5789 | No | ||
| 104 | FOS | 21202 | 14780 | -0.038 | -0.5875 | No | ||
| 105 | PRKCQ | 2873 2831 | 14804 | -0.038 | -0.5883 | No | ||
| 106 | POU2F1 | 5275 3989 4065 4010 | 15053 | -0.048 | -0.6011 | No | ||
| 107 | DUSP8 | 9493 | 15134 | -0.052 | -0.6048 | No | ||
| 108 | CDC25B | 14841 | 15324 | -0.063 | -0.6142 | No | ||
| 109 | TNFRSF1A | 1181 10206 | 15409 | -0.069 | -0.6179 | No | ||
| 110 | ETV1 | 4688 8925 8924 | 15435 | -0.071 | -0.6183 | No | ||
| 111 | IFNG | 19869 | 15440 | -0.071 | -0.6176 | No | ||
| 112 | UBE2D3 | 7253 | 15448 | -0.072 | -0.6170 | No | ||
| 113 | RELB | 17942 | 15563 | -0.083 | -0.6222 | No | ||
| 114 | FGR | 4723 | 15710 | -0.099 | -0.6288 | No | ||
| 115 | MEF2D | 9379 | 15725 | -0.101 | -0.6283 | No | ||
| 116 | UBE2V1 | 12381 | 15795 | -0.111 | -0.6306 | No | ||
| 117 | PPARGC1A | 16533 | 15834 | -0.119 | -0.6311 | No | ||
| 118 | PAK2 | 10310 5875 | 15883 | -0.126 | -0.6321 | Yes | ||
| 119 | EIF4EBP1 | 8891 4661 | 15908 | -0.129 | -0.6317 | Yes | ||
| 120 | NOD2 | 6384 | 15923 | -0.131 | -0.6308 | Yes | ||
| 121 | HCK | 14787 | 15924 | -0.131 | -0.6291 | Yes | ||
| 122 | EGR2 | 8886 | 15952 | -0.137 | -0.6288 | Yes | ||
| 123 | MAP3K8 | 23495 | 15958 | -0.139 | -0.6273 | Yes | ||
| 124 | RIPK1 | 5381 | 15979 | -0.143 | -0.6265 | Yes | ||
| 125 | FKBP8 | 18587 | 16082 | -0.164 | -0.6300 | Yes | ||
| 126 | REL | 9716 | 16118 | -0.172 | -0.6297 | Yes | ||
| 127 | YWHAG | 16339 | 16141 | -0.177 | -0.6286 | Yes | ||
| 128 | MEF2C | 3204 9378 | 16157 | -0.182 | -0.6270 | Yes | ||
| 129 | RPS6KA4 | 23804 | 16167 | -0.184 | -0.6252 | Yes | ||
| 130 | PTPN1 | 5325 | 16237 | -0.200 | -0.6263 | Yes | ||
| 131 | CALM2 | 8681 | 16243 | -0.203 | -0.6240 | Yes | ||
| 132 | LCK | 15746 | 16281 | -0.210 | -0.6233 | Yes | ||
| 133 | CYLD | 18532 | 16309 | -0.218 | -0.6219 | Yes | ||
| 134 | YWHAB | 14744 | 16465 | -0.265 | -0.6269 | Yes | ||
| 135 | MAP2K3 | 20856 | 16576 | -0.300 | -0.6290 | Yes | ||
| 136 | RAC1 | 16302 | 16592 | -0.304 | -0.6259 | Yes | ||
| 137 | IRAK4 | 22379 11185 | 16691 | -0.337 | -0.6269 | Yes | ||
| 138 | RALA | 21541 | 16794 | -0.371 | -0.6277 | Yes | ||
| 139 | MAP2K4 | 20405 | 16842 | -0.384 | -0.6253 | Yes | ||
| 140 | CALM3 | 8682 | 16860 | -0.390 | -0.6212 | Yes | ||
| 141 | MAP2K6 | 20614 1414 | 16879 | -0.397 | -0.6170 | Yes | ||
| 142 | RAF1 | 17035 | 16923 | -0.414 | -0.6140 | Yes | ||
| 143 | MKNK1 | 2504 | 16970 | -0.431 | -0.6110 | Yes | ||
| 144 | IRAK3 | 19617 | 17005 | -0.445 | -0.6071 | Yes | ||
| 145 | PRKCD | 21897 | 17066 | -0.470 | -0.6043 | Yes | ||
| 146 | DDIT3 | 19857 | 17095 | -0.485 | -0.5996 | Yes | ||
| 147 | PRKCA | 20174 | 17118 | -0.497 | -0.5943 | Yes | ||
| 148 | JUNB | 9201 | 17155 | -0.511 | -0.5897 | Yes | ||
| 149 | BIRC2 | 4397 4398 | 17163 | -0.514 | -0.5835 | Yes | ||
| 150 | MAPK14 | 23313 | 17215 | -0.531 | -0.5794 | Yes | ||
| 151 | PIM1 | 23308 | 17294 | -0.572 | -0.5762 | Yes | ||
| 152 | BAD | 24000 | 17315 | -0.582 | -0.5698 | Yes | ||
| 153 | GSK3B | 22761 | 17375 | -0.618 | -0.5651 | Yes | ||
| 154 | MAPK3 | 6458 11170 | 17396 | -0.634 | -0.5580 | Yes | ||
| 155 | MEF2A | 1201 5089 3492 | 17426 | -0.654 | -0.5511 | Yes | ||
| 156 | DUSP10 | 4003 14016 | 17441 | -0.668 | -0.5433 | Yes | ||
| 157 | JUN | 15832 | 17458 | -0.677 | -0.5354 | Yes | ||
| 158 | IL2RA | 4918 | 17525 | -0.716 | -0.5298 | Yes | ||
| 159 | NFATC3 | 5169 | 17539 | -0.726 | -0.5211 | Yes | ||
| 160 | NFKB1 | 15160 | 17596 | -0.757 | -0.5144 | Yes | ||
| 161 | MAP3K14 | 11998 | 17649 | -0.792 | -0.5070 | Yes | ||
| 162 | TOLLIP | 7039 12038 | 17655 | -0.796 | -0.4971 | Yes | ||
| 163 | RELA | 23783 | 17734 | -0.859 | -0.4902 | Yes | ||
| 164 | SYK | 21636 | 17870 | -0.978 | -0.4849 | Yes | ||
| 165 | BCL10 | 15397 | 17878 | -0.984 | -0.4726 | Yes | ||
| 166 | MAP3K7IP2 | 19827 | 17902 | -1.009 | -0.4609 | Yes | ||
| 167 | TRAF6 | 5797 14940 | 17908 | -1.020 | -0.4480 | Yes | ||
| 168 | ITCH | 4928 9185 | 17926 | -1.035 | -0.4356 | Yes | ||
| 169 | LSP1 | 18003 | 17990 | -1.095 | -0.4249 | Yes | ||
| 170 | MAP3K3 | 20626 | 17998 | -1.103 | -0.4111 | Yes | ||
| 171 | CBLB | 5531 22734 | 18038 | -1.159 | -0.3983 | Yes | ||
| 172 | TNFAIP3 | 19810 | 18051 | -1.180 | -0.3837 | Yes | ||
| 173 | DUSP1 | 23061 | 18067 | -1.196 | -0.3691 | Yes | ||
| 174 | IRF4 | 21679 | 18103 | -1.237 | -0.3551 | Yes | ||
| 175 | CREBBP | 22682 8783 | 18124 | -1.271 | -0.3398 | Yes | ||
| 176 | SQSTM1 | 9517 | 18137 | -1.290 | -0.3238 | Yes | ||
| 177 | NFKBIA | 21065 | 18148 | -1.309 | -0.3075 | Yes | ||
| 178 | SLC9A1 | 16053 | 18154 | -1.316 | -0.2908 | Yes | ||
| 179 | CCM2 | 20950 | 18223 | -1.441 | -0.2759 | Yes | ||
| 180 | MAPKAPK2 | 13838 | 18261 | -1.536 | -0.2582 | Yes | ||
| 181 | MAP3K1 | 21348 | 18282 | -1.594 | -0.2387 | Yes | ||
| 182 | MAPK9 | 1233 20903 1383 | 18352 | -1.775 | -0.2196 | Yes | ||
| 183 | BLK | 3205 21791 | 18473 | -2.331 | -0.1961 | Yes | ||
| 184 | NFATC1 | 23398 1999 5167 9455 1985 1957 | 18532 | -2.758 | -0.1637 | Yes | ||
| 185 | EGR1 | 23598 | 18568 | -3.340 | -0.1225 | Yes | ||
| 186 | TP53 | 20822 | 18582 | -3.909 | -0.0729 | Yes | ||
| 187 | RAN | 5356 9691 | 18611 | -5.793 | 0.0003 | Yes |